Talk Overview
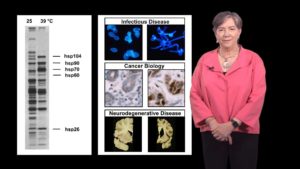
In Part 1, Dr. Lindquist explains the problem of protein folding. Proteins leave the ribosome as long, linear chains of amino acids but they need to fold into complex three dimensional shapes in the extremely crowded environment of the cytoplasm. Since protein misfolding can be disastrous for cells, proteins known as heat shock proteins (HSPs) have evolved to facilitate proper protein folding. Lindquist explains that sometimes the heat shock response becomes unbalanced resulting in human disease. In the case of cancer, HSPs help cancer cells survive many stresses that would typically kill them. In contrast, many neurodegenerative diseases are a result of protein misfolding and aggregation suggesting that, in these diseases, HSPs are not activated when they should be.
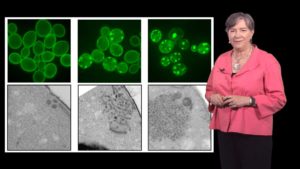
Yeast have many of the same cellular processes as humans including a stress response to aid in protein folding and prevent protein aggregation. In Part 2, Lindquist describes how genetic screens in yeast helped scientists identify mutations that increased the formation of aggregates similar to those found in neurodegenerative diseases. Furthermore a screen in yeast of ~500,000 chemicals identified a number of compounds that prevented protein aggregation. Results from both experiments have since been validated in mice and human neuronal models.
When cells undergo stress, the expression of HSPs increases. In Part 3, Lindquist explains that while most HSPs are expressed only as needed, Hsp90 is expressed in excess. This “buffer” of Hsp90 facilitates the folding of some mutant proteins (such as v-src) that would usually misfold and be degraded by the cell. Thus, Hsp90 potentiates the impact of these mutations. Interestingly, the Hsp90 “buffer” can also act to hide or suppress the impact of other mutations. These “hidden” mutations are found when cells are stressed reducing the pool of available Hsp90. Thus, Hsp90 provides a plausible mechanism for allowing genetic diversity and fluctuating environments to fuel the pace of evolutionary change.
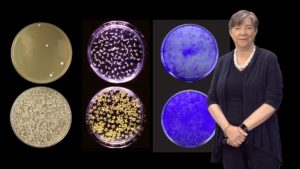
In her last talk, Lindquist focuses on prion proteins. Prions are perhaps best known as the infectious agents in diseases such as mad cow disease. However, Lindquist argues that there are many great things about prions too. They provide a protein-based mechanism of inheritance that allows organisms to develop new traits, quickly and reversibly, and thereby adapt to new environments. Working in yeast, Lindquist and her colleagues were able to identify numerous prion-like proteins that are induced at different levels, depending on the temperature, pH or presence of bacteria. Expression of prions caused heritable, phenotypic changes in the yeast demonstrating that prions are another mechanism by which environmental changes can induce new traits that can be passed onto progeny.
Speaker Bio
Susan Lindquist

Susan Lindquist was a member and former Director of the Whitehead Institute for Biomedical Research. She was also a Howard Hughes Medical Institute Investigator and Professor of Biology at the Massachusetts Institute of Technology. She received her Ph.D. in biology from Harvard and was a postdoctoral fellow of the American Cancer Society. Lindquist was on… Continue Reading




Leave a Reply