Talk Overview
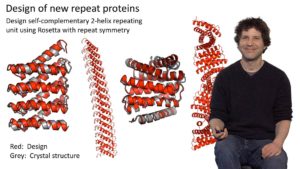
Baker begins his talk by describing two reciprocal research problems. The first is how to predict the 3 dimensional structure of a protein from a specific amino acid sequence, while the second is how to determine the amino acid sequence that will generate a new protein designed to have a specific structure. Baker’s lab is addressing the second of these challenges by developing computer programs (such as Rosetta@Home) that calculate the lowest energy, or most likely, structures for differently folded amino acid sequences. Baker explains how his lab can design a new protein structure, not found in nature, and using the computer programs they have developed, determine the amino acid sequence. It is then possible to back translate to the DNA sequence and synthesize the gene that can then be used to make the protein. When the structures of these synthesized proteins are determined by crystallography and compared to the predicted structures of the designed proteins, they are found to overlap very closely, demonstrating that the protein design algorithms work well.
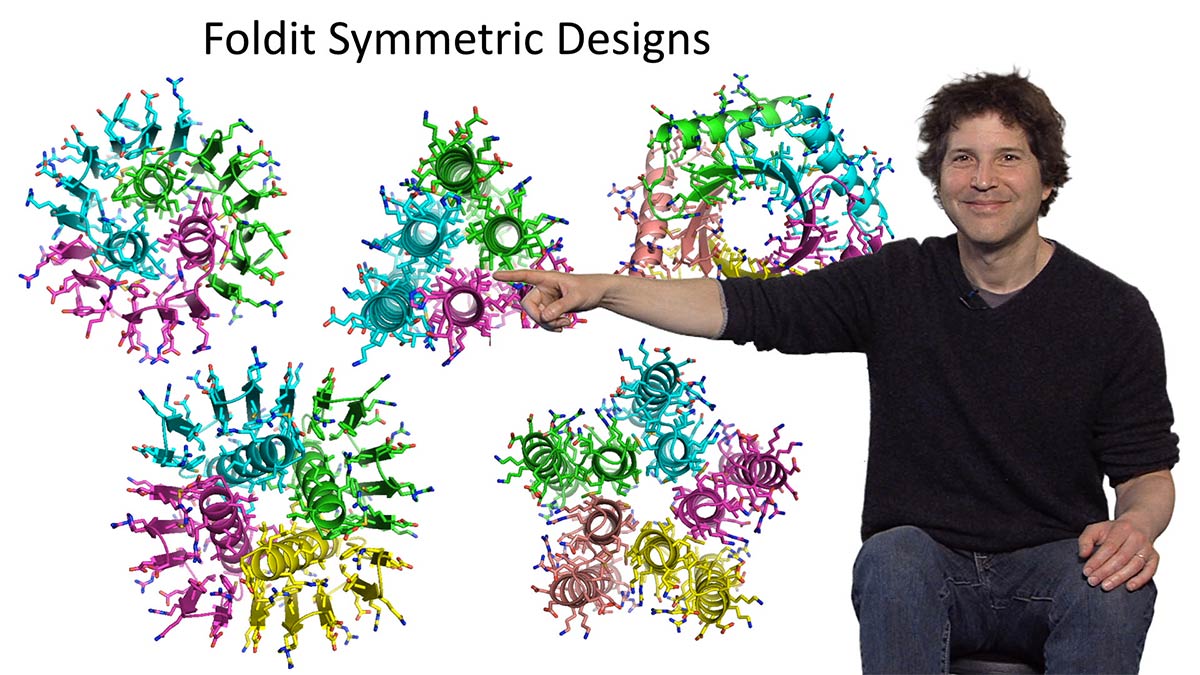
In the second of his talks, Baker tells us how his lab has moved beyond designing new protein structures to designing new protein functions. The first example he describes is the development of an inhibitor of the influenza virus. Baker’s lab designed a protein structure that fits into a highly conserved region of the hemagglutinin protein found on the surface of influenza. Preliminary lab data suggests that this designed protein protects mice from infection with the flu virus. Baker also describes experiments in which proteins were designed to fit together and build multicomponent materials such as nanocages, nanolayers and nanowires.
Speaker Bio
David Baker

David Baker received a BA in Biology from Harvard University and a PhD in Biochemistry from the University of California, Berkeley. Currently, Baker is the Head of the Institute for Protein Design and a Professor of Biochemistry at the University of Washington, and a Howard Hughes Medical Institute Investigator. His research utilizes both experimental and… Continue Reading







Leave a Reply