Talk Overview
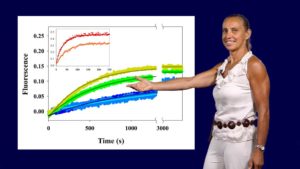
Proteins such as signaling molecules, catalytic enzymes, and membrane transporters are not static but are in a state of constant motion for function. In her first talk, Dorothee Kern explains how she uses techniques such as nuclear magnetic resonance (NMR), X ray crystallography, single molecule FRET and computational simulations to visualize rapid protein dynamics. Kern walks us through the basics of these techniques and explains how they can provide structural and energetic information about a protein and how it functions in the cell.
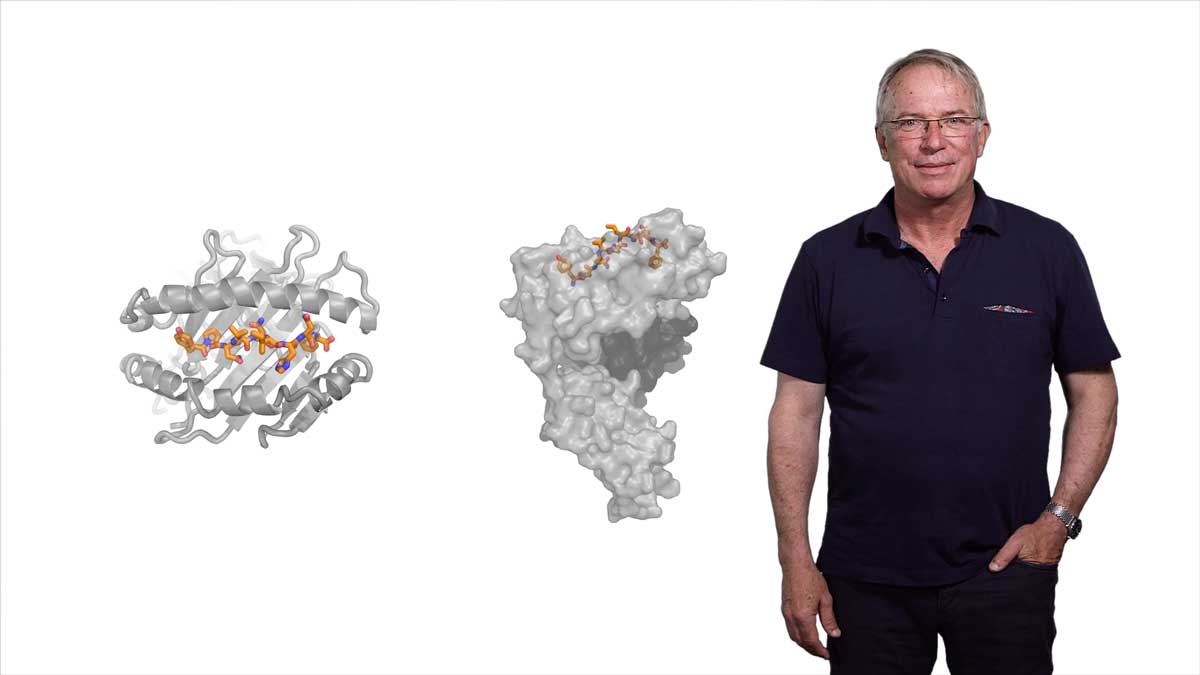
Gleevec (Imatinib) is a very effective drug for treating chronic myelogenous leukemia. It acts by binding to and inhibiting the signaling molecule Abl kinase with extreme specificity. Abl kinase is very closely related to Src kinase, and, in fact, the drug binding pocket for Gleevec is almost identical between the two proteins. Interestingly, however, Gleevec binds to Abl 3000 fold more tightly than it does to Src. Why? In her second talk, Kern answers this question. Her lab used NMR, and the techniques she described in Part 1, to show that the protein dynamics of Abl and Src are dramatically different when Gleevec is bound. In a clever experiment in which they synthesized proteins that were likely evolutionary ancestors of Src and Abl, Kern’s lab was able to show how changes in amino acids throughout the kinases determined the differential binding affinity of Gleevec for Abl kinase over Src kinase. Experiments such as these demonstrate the importance of understanding protein dynamics at the atomic level of the whole protein, not just the drug binding site, when designing new drugs.
Speaker Bio
Dorothee Kern

Dorothee Kern is a Professor of Biochemistry at Brandeis University and an Investigator of the Howard Hughes Medical Institute. Kern strives to understand how proteins function by understanding how they move. She uses biophysical analytical methods, in particular nuclear magnetic resonance (NMR), to analyze protein dynamics. Using NMR, she has followed the high speed motion… Continue Reading







Abi Gartner says
Hello,
I love iBIOLOGY. Thank you for all your hardwork.
You make studying sciences so much more entertaining!
Can I ask – can you provide a transcript for Dorothee Kern’s talks 1/2 on Visualising Protein Dynamics?
Most talks have these, but some don’t.
Thank you