Talk Overview
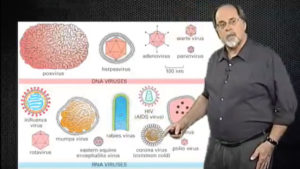
Viruses are extremely simple and small, yet they are responsible for many of the world’s diseases. A virus particle consists of only a genome, a protein coat or capsid, and sometimes a surrounding lipid envelope. To replicate, a virus must successfully enter a host cell, uncoat its genome, and appropriate the host cell machinery to replicate its genome and produce viral proteins. To start off the discussion of viral infection, Part 1 of this lecture will discuss ways in which viruses bind to the surface of host cells. Simian Virus 40, which binds to specific cell surface glycolipids, and Human Papilloma Virus-16, which binds to sites on filoipodia, exemplify different binding mechanisms. Attachment of viruses to the host cell’s plasma membrane activates cell signaling, resulting in endocytosis of the viral particles. This lecture is appropriate for upper level undergraduate and graduate classes studying virology or endocytosis.
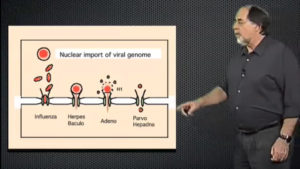
In the second lecture, the next steps in viral infection are described. Endocytosis of plasma membrane bound viruses can occur via a number of mechanisms including caveolar, clathrin, non-clathrin, or lipid raft mediated pathways. The internalized virus is enclosed in an endosome that may undergo increasing acidification, resulting in acid mediated fusion between the viral envelope and the vesicle membrane. Following membrane penetration, the virus, once again, makes use of cellular machinery such as microtubules and their motors, to transfer its genome to the nucleus. Helenius describes experiments from his lab and others that have deciphered these complex processes.
Part 3 focuses on a single virus, the Vaccinia virus, as a model for cell binding, signaling and endocytosis. Fluorescently labeled Vaccinia viruses bind to and surf along host cell filopodia. Helenius lab members noticed that when Vaccinia, unlike other viruses, reached the surface of the cell body it caused the plasma membrane to form blebs. Further experiments showed that the virus tricks the cell into thinking it is apoptotic debris. This induces blebbing and subsequent uptake of the virus by macropinocytosis. Additionally, automated high throughput siRNA screening was used to screen a large number of infected cells for host genes required for Vaccinia virus uptake. Analysis of the genes identified allowed host factors and processes critical to viral infection to be identified. Expansion of this technique may provide a new source of information on pathogen-host interactions.
Speaker Bio
Ari Helenius

Ari Helenius is from Finland where he studied Biochemistry in the University of Helsinki. During his PhD thesis work, which focussed on the analysis of membrane proteins, he and his colleagues started to work on the biochemical properties of enveloped animal viruses. The simplicity and elegance of Semliki Forest virus and its membrane proved to… Continue Reading









Leave a Reply